Curse of Dimensionality
Learn about the ‘Curse of Dimensionality’ and its impact through this article.
The Curse of Dimensionality sounds like it’s something straight out of the wizarding world, but it really is only a very common term you’ll come across in the Machine Learning and Big Data world. This term describes how the increase in input data dimensions results in an exponential increase in computational expense and efforts required for processing and analyzing that data.
In this article, we will cover the following sections:
What is the Curse of Dimensionality?
The curse of dimensionality basically refers to the difficulties a machine learning algorithm faces when working with data in the higher dimensions, that did not exist in the lower dimensions. This happens because when you add dimensions (features), the minimum data requirements also increase rapidly.
This means, that as the number of features (columns) increases, you need an exponentially growing number of samples (rows) to have all combinations of feature values well-represented in our sample.
With the increase in the data dimensions, your model –
- would also increase in complexity.
- would become increasingly dependent on the data it is being trained on.
This leads to overfitting of the model, so even though the model performs really well on training data, it fails drastically on any real data.
Quite a few algorithms work only on tall, svelte datasets with fewer features and more samples. Hence, to remove the curse afflicting your model, you might need to put your data on a diet – i.e., reduce its dimensions through feature selection and feature engineering techniques. Let’s see how this is done!
Explore machine learning courses
Best-suited Machine Learning courses for you
Learn Machine Learning with these high-rated online courses
Dimensionality Reduction to the Rescue
What you need to understand first is that data features are usually correlated. Hence, the higher dimensional data is dominated by a rather small number of features. If we can find a subset of the superfeatures that can represent the information just as well as the original dataset, we can remove the curse of dimensionality!
This is what dimensionality reduction is – a process of reducing the dimension of your data to a few principal features.
Fewer input dimensions often correspond to a simpler model, referred to as its degrees of freedom. A model with larger degrees of freedom is more prone to overfitting. So, it is desirable to have more generalized models, and input data with fewer features.
Why is Dimensionality Reduction necessary?
- Avoids overfitting – the lesser assumptions a model makes, the simpler it will be.
- Easier computation – the lesser the dimensions, the faster the model trains.
- Improved model performance – removes redundant features and noise, lesser misleading data improves model accuracy.
- Lower dimensional data requires less storage space.
- Lower dimensional data can work with other algorithms that were unfit for larger dimensions.
How is Dimensionality Reduction done?
Several techniques can be employed for dimensionality reduction depending on the problem and the data. These techniques are divided into two broad categories:
Feature Selection: Choosing the most important features from the data
Feature Extraction: Combining features to create new superfeatures.
Now, we are going to demonstrate how to get rid of the curse of dimensionality. We will be performing dimensionality reduction through a common linear method – Principal Component Analysis (PCA).

Demo: Implementing PCA for Dimensionality Reduction
Problem Statement:
A large number of input dimensions can cause a model to slow down during execution. So, we perform Principal Component Analysis (PCA) on the model to speed up the fitting of the ML algorithm.
PCA projects data in the direction of increasing variance. The features having the highest variance are the principal components. Let’s see how to implement PCA using Python:
Dataset Description:
We will be using the iris flower dataset for this.
This dataset has 4 features:
- sepal_length – Sepal length in centimeters
- sepal_width – Sepal width in centimeters
- petal_length – Petal length in centimeters
- petal_width – Petal width in centimeters
- species – Species of iris
The ‘species’ column is our target variable
Tasks to be performed:
- Loading the dataset
- Standardizing the data onto a unit scale
- Projecting PCA to two-dimensions
- Concatenating the Principal Components with the target variable
- Visualizing the 2D Projection
Step 1 – Loading the dataset
import pandas as pd
iris = pd.read_csv(‘IRIS.csv’)
iris.head()
Step 2 – Standardizing the data onto a unit scale
Before we apply PCA, the features in the given data need to be standardized onto a unit scale (mean = 0 and variance = 1). This ensures the optimal performance of many ML algorithms. For this, we use StandardScaler, as shown:
from sklearn.preprocessing import StandardScaler
features = [‘sepal_length’,’sepal_width’,’petal_length’,’petal_width’]
#Separating the features
x = iris.loc[:, features].values
#Separating the target variable
y = iris.loc[:,[‘species’]].values
#Standardizing the features
x = StandardScaler().fit_transform(x)
Step 3 – Projecting PCA to 2D
The initial dataset has 4 features: sepal length, sepal width, petal length, and petal width.
Now, we will be projecting this data into two principal components – PC1 and PC2. These will now be our main dimensions of variance:
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
components = pca.fit_transform(x)
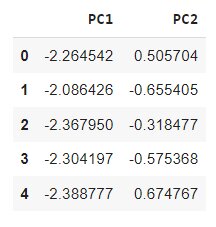
data = pd.DataFrame(data = components,
columns = [‘PC1’, ‘PC2’])
data.head()
Step 4 – Concatenating the Principal Components with the target variable
Now, we will concatenate the DataFrame, created in the above step, along axis = 1.
final_data = pd.concat([data, iris[[‘species’]]], axis = 1)
final_data.head()
Thus, we have performed dimensionality reduction and removed the curse of dimensionality that had afflicted our data, by performing PCA on it.
Step 5 – Visualizing the 2D Projection
import matplotlib.pyplot as plt
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel(‘Principal Component 1’, fontsize = 15)
ax.set_ylabel(‘Principal Component 2’, fontsize = 15)
ax.set_title(‘2 component PCA’, fontsize = 20)
targets = [‘Iris-setosa’, ‘Iris-versicolor’, ‘Iris-virginica’]
colors = [‘red’, ‘green’, ‘orange’]
for target, color in zip(targets,colors):
indicesToKeep = final_data[‘species’] == target
ax.scatter(final_data.loc[indicesToKeep, ‘PC1’]
, final_data.loc[indicesToKeep, ‘PC2’]
, c = color
, s = 50)
ax.legend(targets)
ax.grid()
As demonstrated in the graph above, we can see well separated the classes are from each other.
Endnotes
Dimensionality Reduction is an important yet often overlooked step in the ML workflow of organizations in general. In a domain where more data is considered to be a good thing, we have rediscovered how low-quality unnecessary data can create more problems than before.
Hope this article helped you understand the importance of keeping the features of data to a minimum. Artificial Intelligence & Machine Learning is an increasingly growing domain that has hugely impacted big businesses worldwide. Interested in being a part of this frenzy? Explore related articles here.